The Hannee software platform (Heidelberg Analog Neural Network Evolution Environment) was developed to control, train, and test our hardware neural networks. It combines low-level hardware access modules with training and evaluation algorithms and two user interfaces, in order to provide control of the hardware on an abstract, though efficient, level.
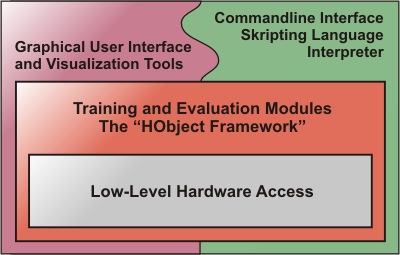 Structure of the Hannee software platform
Structure of the Hannee software platform
The figure shows a schematic of the software structure. On top of the hardware interface classes are the training and evaluation modules. This part of the program does most of the actual work. The program can be driven by a graphical user interface which provides additional visualization tools or, optionally, by a commandline interpreter.
Main features of the software include:
Please enjoy our gallery of Hannee screenshots. The various images show the main parts of the user interface in action.
The Hannee Main Window. Here, all available modules are shown and can be selected.
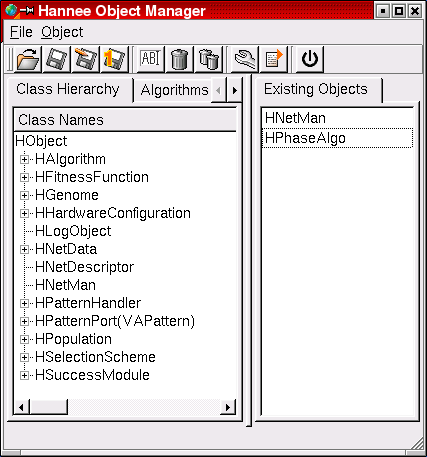
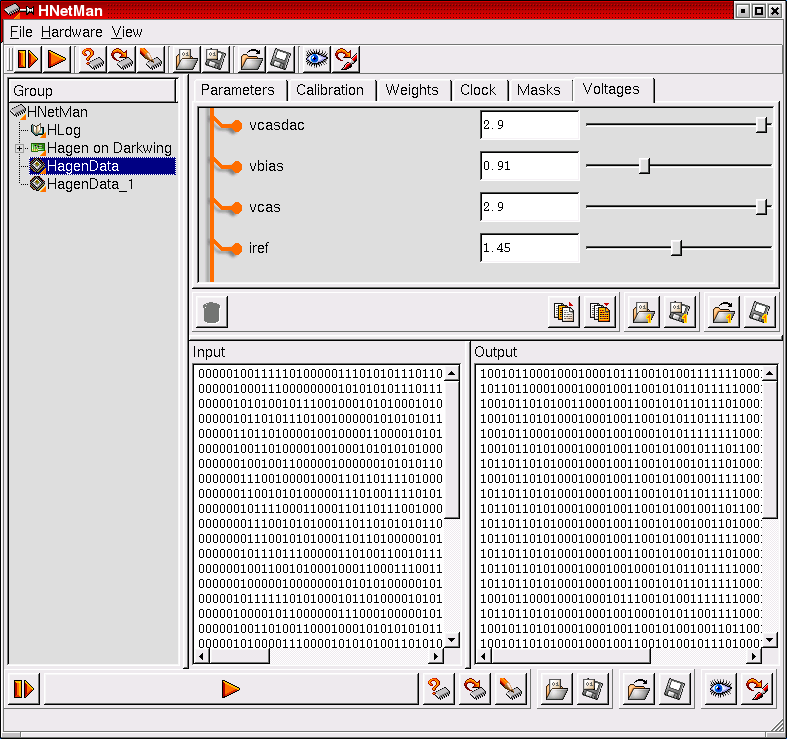
The HNetMan object completely encapsulates the hardware access. Its own modular structure allows to control arbitrary hardware configurations and to host complex network descriptions to be implemented on the chip. In particular, it can be used to monitor the current state of the hardware as well as to manipulate the network configuration online.
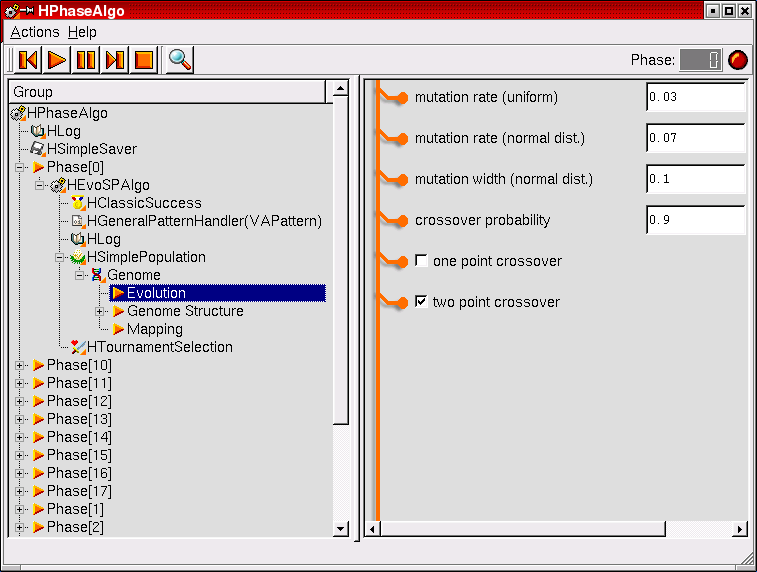
A nice example for an algorithm module. The HPhaseAlgo is divided into various phases that each train different parts of the network. Within each phase, a different evolutionary algorithm can be used which can be configured with the desired parameter settings. The user interface allows a comfortable navigation through the specific algorithm parameters.
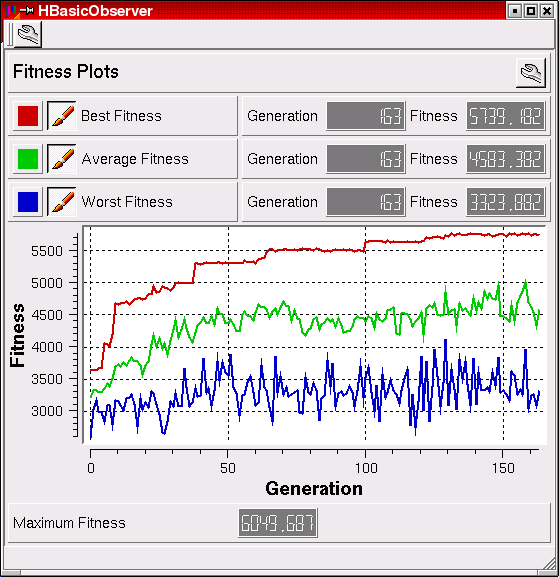
The HBasicObserver is a tool to visualize the progress of any evolutionary algorithm within the Hannee framework. The "fitness" is a measure for how well the network solves the given task. At first, the network knows nothing (low fitness), but in the course of the evolution, the fitness increases and might eventually approach the maximum possible value.
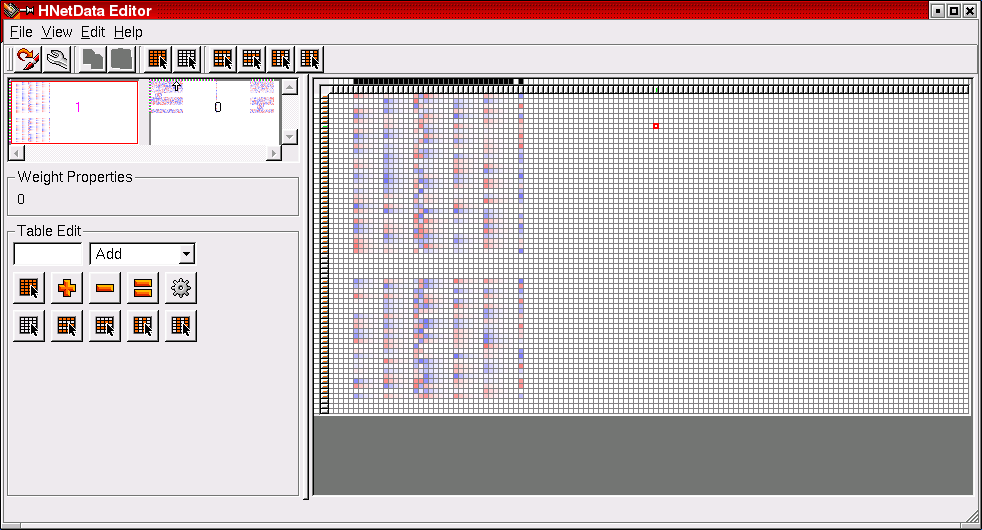
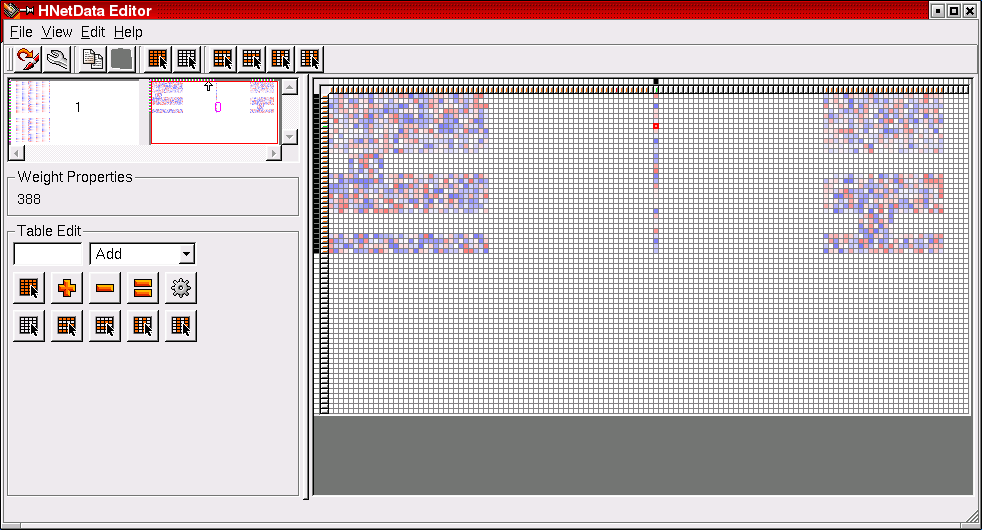
The HNetDataEditor provides comfortable means to construct network descriptions in an interactive mode. Positive weights are a shade of blue, negative weights are red. This example shows a network which is distributed among two blocks. The first block is enlarged in the right half of the HNetDataEditor window.
This image shows the second block of the example network above as it appears in the HNetDataEditor window. This network is used for the well-known benchmark problem of predicting the cellular localization sites of E.coli proteins. Click here to read more about this problem and how it can be solved by evolutionary algorithms using the Hannee software.
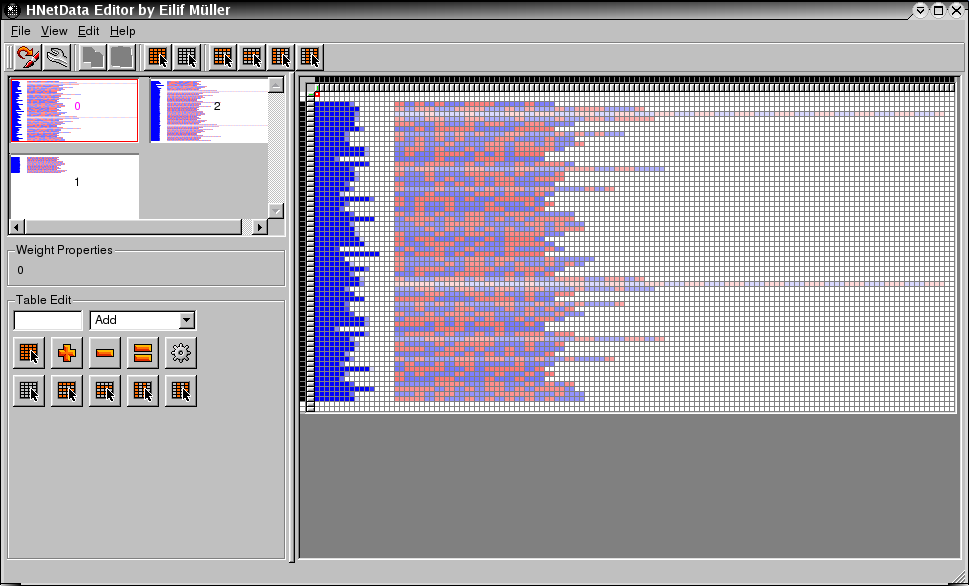
This network has been trained for the recognition of patterns in biological sequences. Click here to learn more about this interesting application of our hardware neural network training environment.
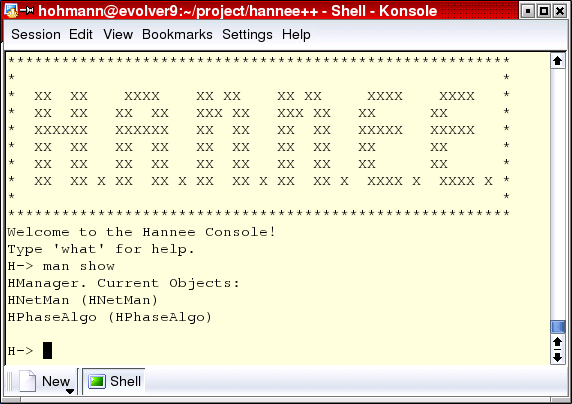
The commandline interface allows to use the complete functionality of the Hannee software without any graphical user interface. This is particularly useful to perform complex experiments (e.g. during the weekend) by automatically executing pre-defined scripts.
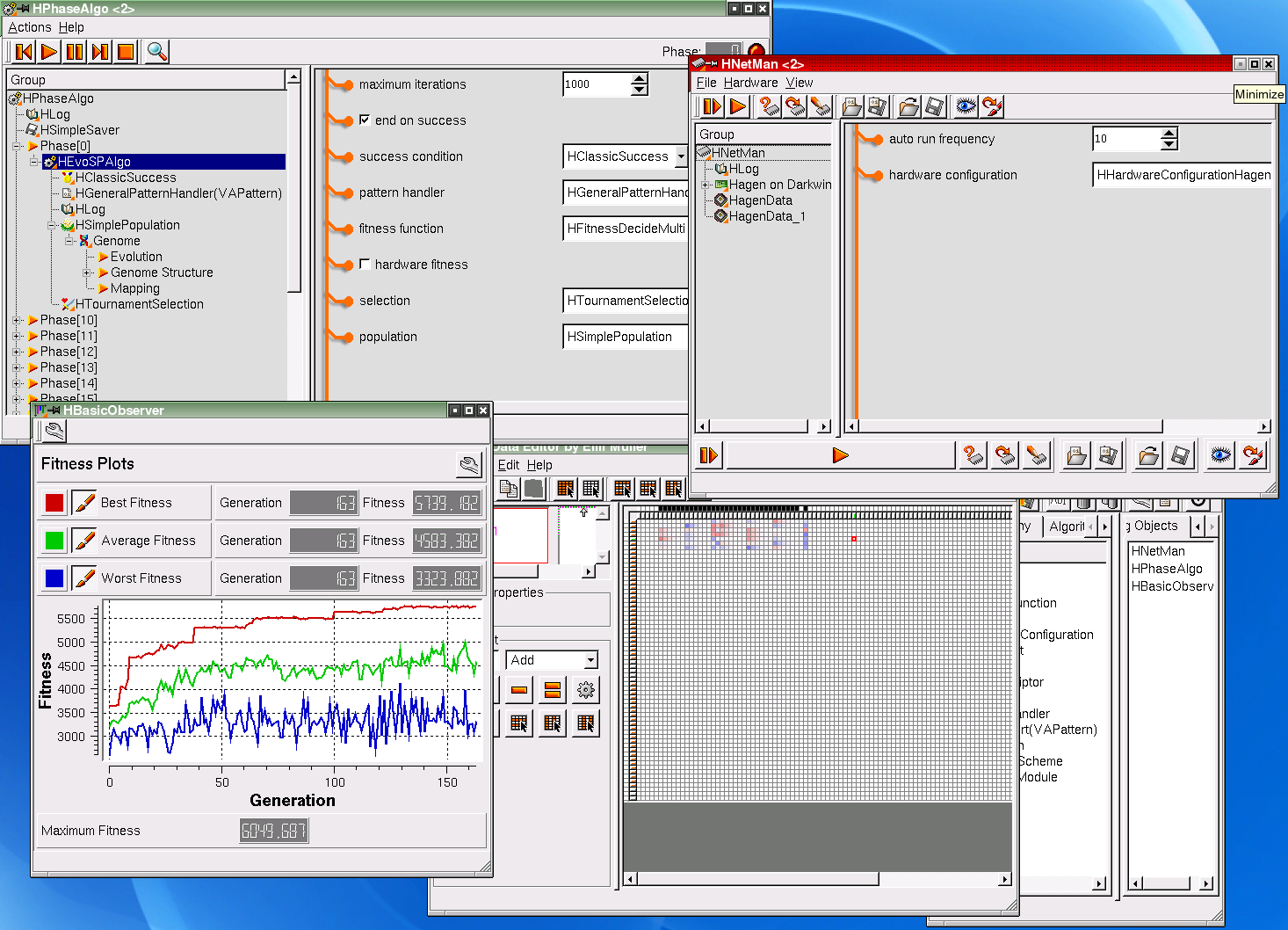
This image shows a complete screenshot of the Hannee software in action.
Electronic Visions Group – Prof. Dr. Johannes Schemmel
Im Neuenheimer Feld 225a
69120 Heidelberg
Germany
phone: +49 6221 549849
fax: +49 6221 549839
email: schemmel(at)kip.uni-heidelberg.de
(All applications only via 'Open Positions')
How to find us